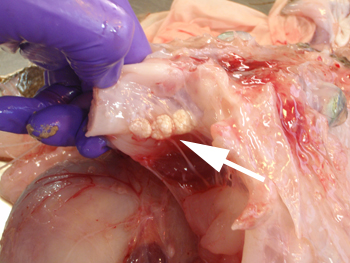
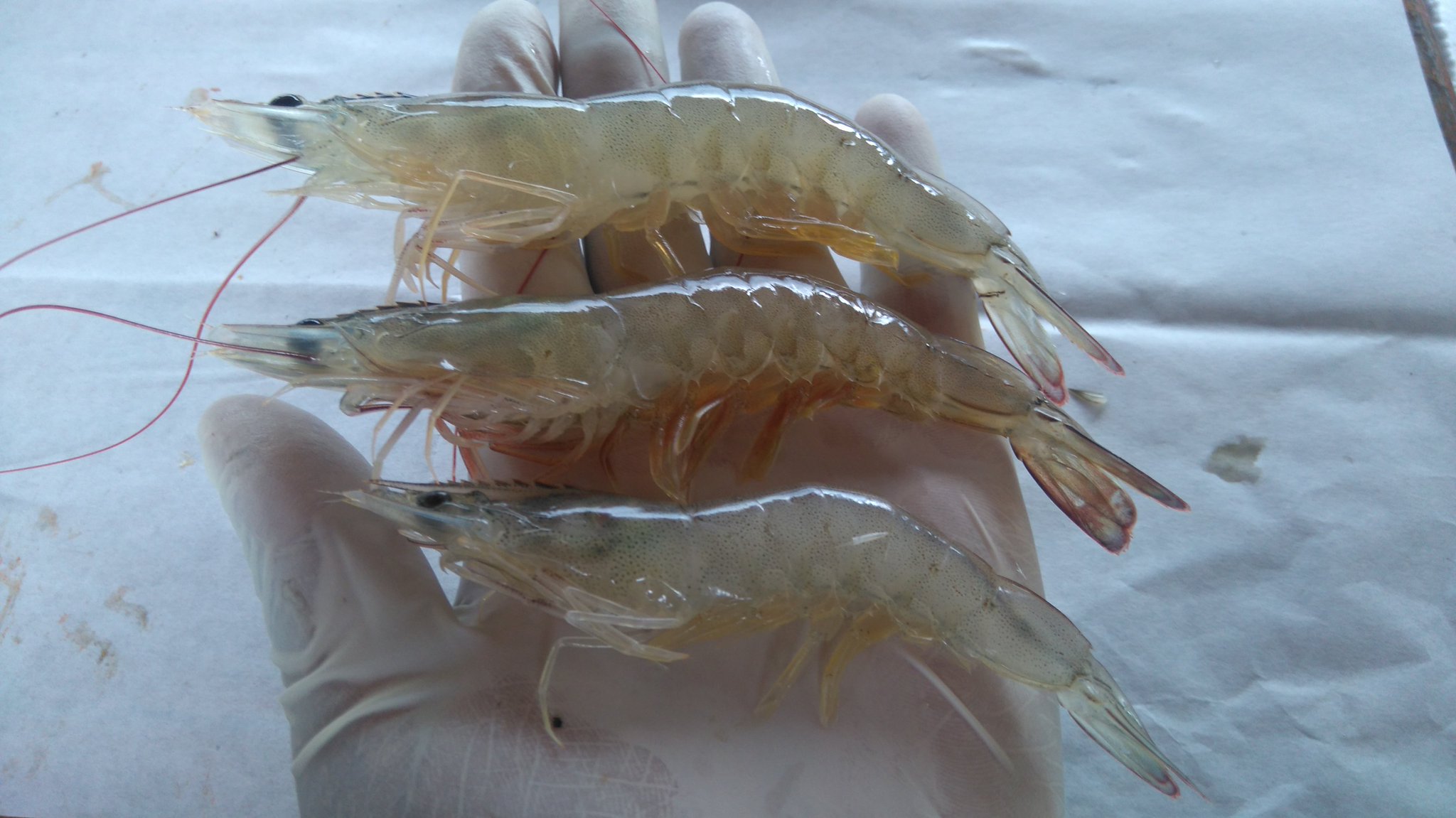
Crustacean pathogens
Microsporidia a becoming important pathogens of crustacea including farmed shrimp and crabs. We have sequenced the genomes of several pathogens in the Enterocytozooniade family. These are interesting from an evolutionary perspective as the whole family appears to have lost the electron transport chain and glycolysis and thus have no obvious intrinsic means of energy generation.

Encephalitozoon cuniculi
E. cuniculi is a mammal-infecting microsporidian for which the whole genome sequence is available and can be cultured in mammalian tissue culture systems which makes it a good system to work with in the lab.
Within host cells. large numbers of spores develop within a parasitophorous vesicle.
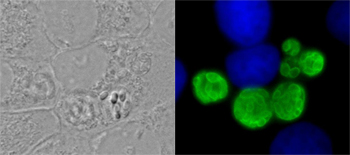
Diversity in the environment
The study of biodiversity has been revolutionised by the molecular era, where high throughput molecular methods can screen environments for new diversity on unprecedented scales. Crucially, molecular methods allow easy access to that microbial diversity not visible to the naked eye. However, one trophic group that are rarely targeted using this strategy are the parasites.
Parasites are important regulators of population size, help define community structures, and a likely a major component of biological diversity. Importantly they also can represent a risk for new disease and infection in wildlife and humans, yet are largely overlooked in terms of biodiversity sampling.
We are carrying out a study of parasite diversity in the environment by surveying the diversity of microsporidia parasites using a high-throughput 454 sequencing approach.

Spraguea lophii
Microsporidia are difficult to culture in large numbers and without a genetic manipulation system, protein characterisation in this group is challenging.
Spraguea lophii is a monkfish-infecting microsporidian which forms large cysts of spores in enlarges fish cells called xenomas.
These are abundantly available from environmental samples, which makes it a good model system for genomic, proteomic and metabolomic research.